Marketing Materials
iReceptor Plus Brochure
The iReceptor Plus brochure presents how this project can unleash the power of the Adaptive Immune System to treat disease via personalized immunotherapy.
iReceptor Plus provide a queriable set of AIRR-seq data repositories following FAIR (findable, accessible, interoperable, and reusable) principles and the AIRR Community’s data standards for sharing AIRR-seq data.
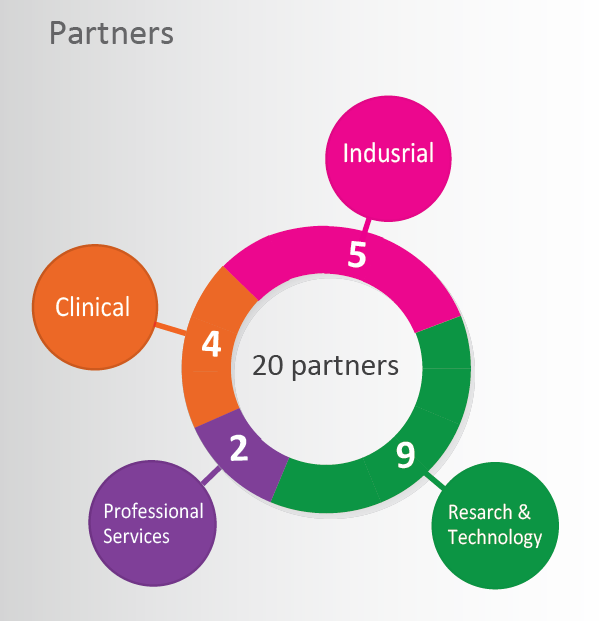
The iReceptor Plus Project by Numbers
The European Union (EU) and the Canadian government have awarded €8.4 million to the international iReceptor Plus Consortium, which is composed of more than 20 partners from 9 countries to promote human immunological data storage, integration and controlled sharing for a wide range of clinical and scientific purposes.
iReceptor Plus Fact Sheet
The iReceptor Plus fact sheet provides all the required information about the project, which is co-funded by the EU and Canada.
The fact sheet contains also information about the six other projects included at the EUCAN cluster.