Blog
The “Art Gallery” of Data: How the AIRR Community is Making Health and Genomics Data More Valuable through Sharing
04/02/2019
By Judy Siegel-Itzkovich
If two heads are better than one, then numerous groups of researchers working in numerous countries around the world but sharing data aimed at the same target are preferable to independent and isolated scientific work.
The international iReceptor Plus consortium – a four-year project financed jointly by the European Union and the government of Canada – is devoted to collecting big data on the immune systems of healthy individuals and patients that can be shared for a wide range of scientific and clinical uses. The sequence data will be stored in databanks in a number of countries; iReceptor Plus will speed up the analysis and international sharing of these data among labs, diseases and institutions.
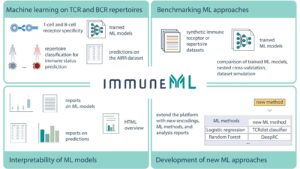
The main focus of this consortium is to analyze and share antibody /B-cell and T-cell receptor sequences more efficiently. High-throughput sequencing of B and T cell receptors is routinely being applied in studies of adaptive immunity.

The work of the iReceptor Plus Consortium is based on protocols and standards developed by a community initiative called the AIRR Community. Launched in 2015, the Adaptive Immune Receptor Repertoire (AIRR) Community is a community-driven organization that is organizing and coordinating stakeholders in the use of next-generation sequencing (NGS) datasets to study medicine
Sequencing is the process of determining the order of four nucleotides (bases) – adenine, guanine, cytosine and thymine – in DNA. The development of rapid NGS DNA sequencing methods has greatly speeded up biological and medical research and discovery. With its ultra-high throughput, scalability and speed, NGS has revolutionized the biological sciences, enabling researchers to perform many types of applications and study biological systems at a level never before possible. Today’s complex genomic research questions demand a depth of information beyond the capacity of traditional DNA sequencing technologies. NGS has filled that gap and become an everyday research tool to address these issues.
NGS applied to antibody-B-cell and T-cell repertoires produces AIRR-seq data. AIRR sequencing offers enormous potential for understanding the dynamics of the immune repertoire in vaccinology, infectious disease, autoimmunity and cancer biology. Since analyzing and sharing these data is not an easy task, the AIRR Community is developing established standards for the production, sharing, storage and analysis of antibody/B-cell and T-cell receptor repertoire sequence (AIRR-seq) data derived from high-throughput sequencing of B cells, plasma cells and T cells.

The AIRR Community consists of 150 members – academic and industrial researchers, including basic-science and clinical immunologists, protein-immunotherapeutics engineers, statisticians, bioinformaticians, computer-security experts, and scholars in the relevant aspects of ethics, law, and policy.
Six Working Groups have been organized to develop recommendations for a common repository for AIRR-Seq data; minimal standards for publishing or storing AIRR sequence data, resources and guidelines for the evaluation of molecular and statistical methods for AIRR sequence data, data standards for software development, AIRR-Seq data representation; and rules and guidelines for inferring germline sequences from AIRR-Seq data.
The Community has already met once in Vancouver in 2015 and then in 2016 and 2017 the Community Meetings convened in Bethesda, Maryland and were supported by the US National Institutes of Health (NIH).
The next scheduled Community Meeting will be in May at the University of Genoa in Italy. The theme of this meeting, “Bridging the Gaps,” will discuss technological gaps between the amounts of accumulated data and the ability to process them, as well as the need for more involvement of stakeholder communities (industry, clinicians, patient communities) to raise the standards developed by the Community. Among the confirmed speakers are Dr. Sai Reddy from ETH Zurich and Dr. Antonio Lanzavecchia from the Institute for Research in Biomedicine (IRB Bellinzona).
Additional resources
Click here for agenda and registration information or send an email to meetings@AIRRC.antibodysociety.org