Blog
AIRR-seq can teach about fundamental immune processes, with broad implications for biomed
30/01/2022
By Judy Siegel-Itzkovich
The immune system’s success in fighting countless evolving pathogens depends on a dynamic and diverse set of B and T cell receptors (BCRs and TCRs).
The diversity and longevity of immunological memory provide detailed insights into the past and present encounters of the human immune system. It can teach us about fundamental immune processes and reveal dysregulation, with broad implications for biomedicine.
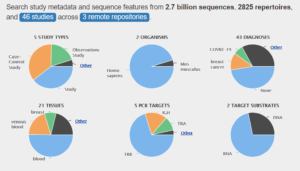
TCR and BCR repertoires, or adaptive immune receptor repertoires (AIRRs), are the front line of adaptive immunity. AIRR-seq is becoming a common tool to study immune system dynamics.
Understanding the genetic factors that influence the composition and dynamics of these repertoires is of major scientific and clinical importance. The chromosomal loci encoding for the variable regions of TCRs and BCRs are challenging to decipher because of repetitive elements and undocumented structural variants.
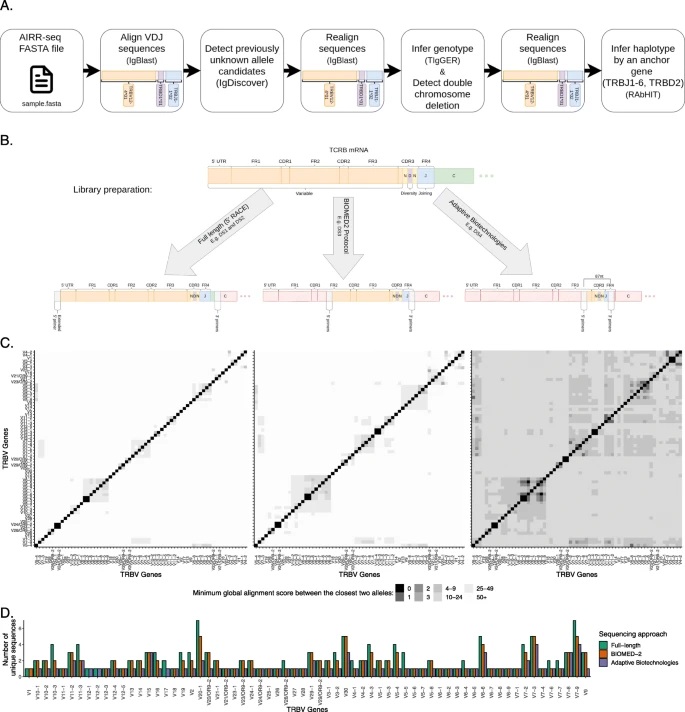
This is one of the challenges confronted by a new research paper entitled “T cell receptor beta germline variability is revealed by inference from repertoire data”, which was published in Genome Medicine. The article was published as part of the iReceptor Plus Project by Aviv Omer, Ayelet Peres, Pazit Polak, Gur Yaari, Oscar Rodriguez, William Lees, and colleagues.
The authors adapted AIRR-seq-based methods that were recently developed for B cells, to T cells. These methods enable genotype and haplotype inference and discovery of undocumented alleles, circumventing the need for direct genomic sequencing.
A major obstacle was that this approach relies on complete coverage of the receptors’ variable regions, which is suitable for B cells, but most T cell studies sequence only a small fraction of that region.
The researchers therefore had to adapt the B cell pipeline for full and partial AIRR-seq TCR data sets. The pipeline also deals with gene assignment ambiguities, which is especially important in the analysis of data sets of partial sequences.
“We adapted BCR inference methods to TCRs and discover a plethora of unknown allelic variations in the coding and non-coding regions. We also discovered how a hidden single nucleotide in the TRBD gene is associated with dramatic changes in the expressed repertoire,” the team concluded. “Our findings provide a basis for annotation of TCR repertoires for future basic and clinical studies.”
“It is not trivial to spot such tiny differences in the TCR genes, but it appears to make huge changes in the repertoire that until now were overlooked,” noted Yaari, who is the coordinator of the iReceptor Plus Project.