Blog
Machine learning technique helps distinguish between healthy and cancerous tissue
27/06/2019
By Judy Siegel-Itzkovich
A group of researchers at the University of Texas Southwestern Medical Center in Dallas Texas, which is part of the iReceptor Plus Project, have published an article in the prestigious journal Cancer Research. The article focuses on using a machine- learning approach that can differentiate between normal and cancerous tissue by looking at the immune repertoire.
The title of the main article was “Biophysicochemical Motifs in T-cell Receptor Sequences Distinguish Repertoires from Tumor-Infiltrating Lymphocyte and Adjacent Healthy Tissue,” and the researchers were Jared Ostmeyer, Scott Christley, Inimary Toby, and Lindsay Cowell.
In addition, the journal published a research highlight article that discusses some of the significant results.
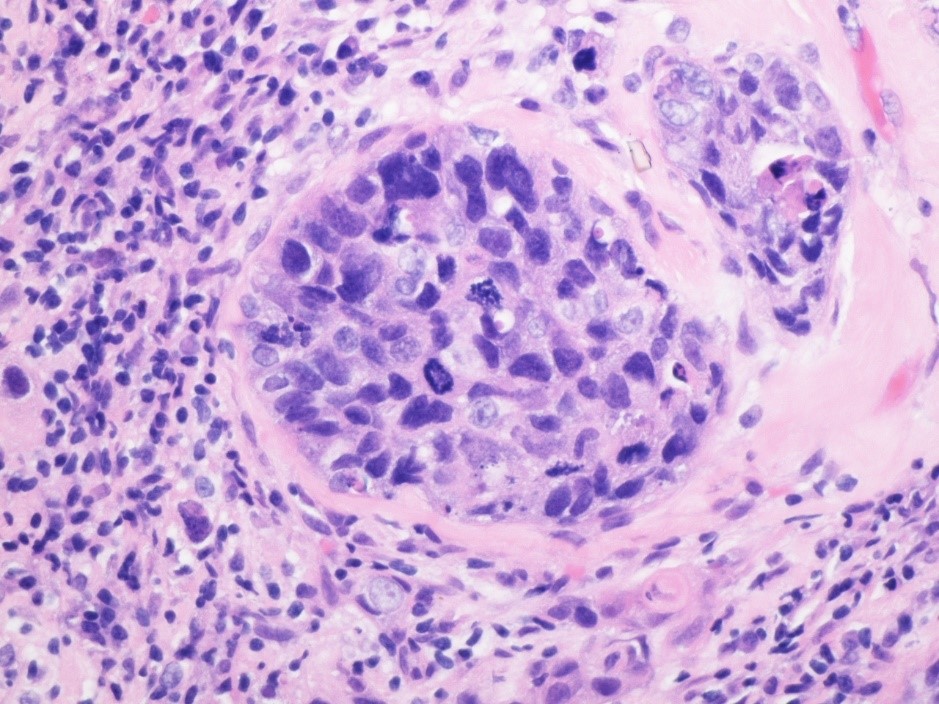
Dr. Ostmeyer, a postdoctoral fellow at UTSW who works with Dr. Cowell, spoke in an interview in detail about his work on the research team.
Born in Joplin, Missouri to a journalist and a social worker, Ostmeyer studied physics and mathematics at the University of Arkansas, did his doctorate at the University of Chicago, and for the past three years has been doing research at the huge Texas academic medical center.
“The part of the immune system we care about is the adaptive immune system, which is composed of B and T cells; each cell type expresses a different immune receptor. There is a random process in the body in which an immune cell creates immune receptor genes with different properties,” he said. “It might bind to a flu virus or a protein expressing a cancer antigen. If you are lucky, you have the right one to bind to a disease antigen. We can now sequence large numbers of immune receptor genes from a patient.”
Ostmeyer explained that his team uses next-generation DNA sequencing techniques with “special tricks to sequence the regions that contain the immune receptor genes”. Because DNA sequencing keeps dropping in cost, it is now about $500 dollars per person, but the cost will continue to fall.
Once they sequenced all relevant genes, they obtained traces of the immune-response history. Then we interpret the results,” said Ostmeyer, who spends all his work time in front of a computer screen.
The UTSW team used an approach they developed for better and more quickly diagnosing multiple sclerosis and applied it to cancer – specifically colorectal and breast cancer. Immune repertoire deep sequencing allows comprehensive characterization of the antigen receptor–encoding genes in a lymphocyte population.
“We hypothesized that this method could enable a novel approach to diagnose disease by identifying antigen receptor sequence patterns associated with clinical phenotypes. In this study, we developed statistical classifiers of T-cell receptor repertoires that distinguish tumor tissue from patient-matched healthy tissue of the same organ,” the team wrote. “This study presents a novel computational approach to identify T-cell repertoire differences between normal and tumor tissue.”
“It was surprising that it worked, but we found an immune receptor pattern only present in tissue from colorectal tumor but not healthy colon tissue. We are also working on ovarian cancer but have not yet published the results. If these diseases are diagnosed early, which our technique may make possible, it is much more treatable,” Ostmeyer stressed.
“We’ve developed a way to relate immune receptors in the patient to disease states. We look for similar immune receptors in patients with the same disease and check that the similar ones are not found in patients without the disease.”
The iReceptor Plus Project, he continued, is important because it makes data available. “My job is interpreting the data. There are other datasets to diagnose viruses or any disease that has an immune response,” Ostmeyer concluded. “If the immune system is involved, there is an opportunity to see if there are immune receptors to diagnose the disease. The immune system is very rich with information. We study patient histories at the molecular level.”
Ostmeyer added that the methods the team developed to relate immune receptor sequences to a patient’s disease states are special in one very important way. “We have shown that when we get new patients, the patterns that we have found in the immune receptor sequences, which we use to identify a disease state, still hold true, albeit at a slightly lower level of performance. In other words, what we’ve found appears to be statistically significant.”
It’s “easy to trick yourself in my line of work. If you toss a coin four times, there are 2^4 = 16 possible outcomes. If we try 16 different models to predict the coin toss, one of those 16 models will predict the coin tosses. It doesn’t mean the model is statistically significant. The trick is to see if the model still works if you toss the coin four more times. That’s what we’ve done, and our predictions still work. With each paper, results get more exciting.”