Blog
Desperation for effective vaccines highlights the importance of projects such as iReceptor Plus for breakthrough in vaccine research
19/03/2020
By Judy Siegel-Itzkovich
The world’s people and governments would give anything for an effective vaccine that could put a halt to the current frightening and devastating COVID-19 epidemic.
Since Edward Jenner introduced the first successful vaccine against smallpox in 1796 – investigating his observation that milkmaids who had previously been infected with cowpox did not later catch smallpox – vaccination has been a fundamental pillar of human health.
Nevertheless, our understanding of how the immune system (or specifically, immune receptor repertoires) recognizes pathogens (viruses, bacteria and so on) has remained rather limited so far. Many research groups believe that to solve this problem, we need to combine computational biological, machine learning and high-throughput and large-scale immunology datasets.
Associate Prof. Victor Greiff at the Laboratory for Computational and Systems Immunology in the University of Oslo and his team have summarized, for the general audience, the state of the art of knowledge of combining computer science and immunology for vaccine research.
Projects such as iReceptor Plus are key to solving current problems in vaccine research as they provide the basis for mining adaptive immune receptors across many different datasets from healthy and vaccinated individuals, Greiff stated.
“This will help us understand how the adaptive immune response reacts to vaccination and how we need to engineer vaccines to elicit broader and more efficient vaccination strategies.”
Greiff started his research group at the University of Oslo in 2018. The team members develop computational and experimental tools to decipher, read, predict and re-engineer antibody and T-cell repertoires so as to develop fundamentally novel immunodiagnostics, vaccines and immunotherapeutics. He uses his time away from science to catch up on current immunology topics, technology developments, and state of the art algorithms.
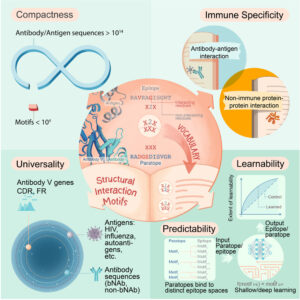
His team recently published an article on the issue entitled Vaccine research: In need of a Rosetta stone. The article, which was published in Vaccine Blog states that vaccination rests on the ability of the immune system to form “immunological memory.” – the immune system remembers successful battles of the past to mount more efficient immune responses upon meeting the pathogen a second time. Just like our brain remembers many things from our childhood up to our present age, our immune system keeps a record of every infection it has encountered over the lifetime.
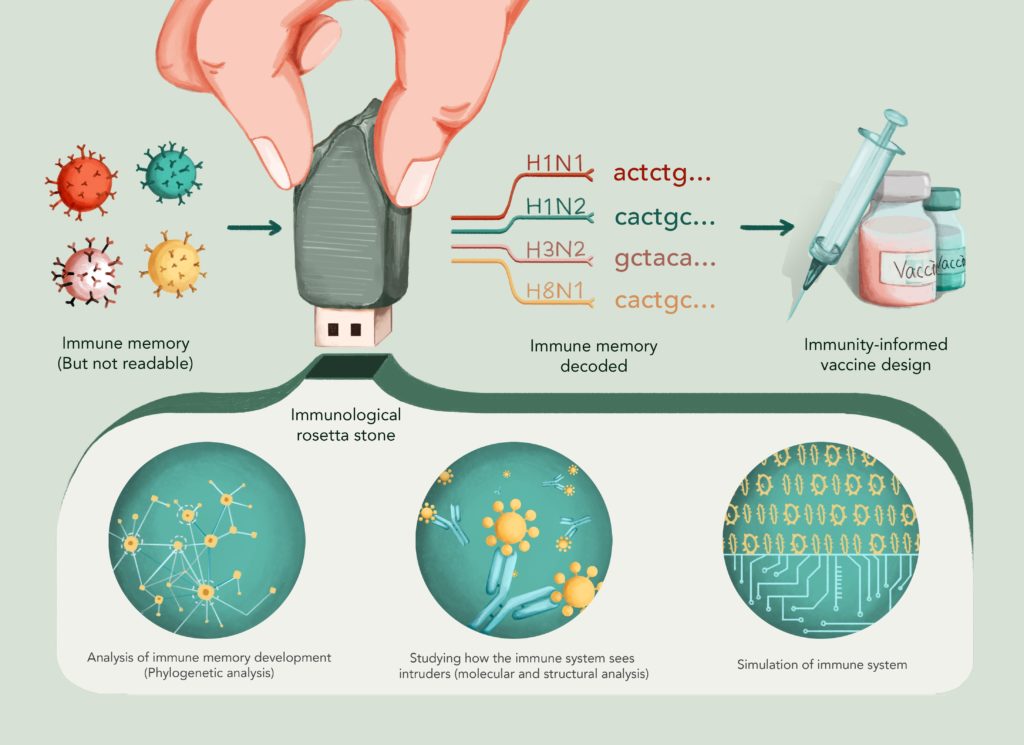
Immune memory is recorded in the most universal USB drive of all – the DNA. Most immune cells (or “adaptive” immune cells) have unique sections of DNA and are each programmed to recognize different targets. Our immune memory is a combination of those soldiers that the body has selected and multiplied in previous fights. A successful vaccine needs to arouse enough soldiers and train them to recognize a specific target.
Over the past 10 years, it has become possible to read the DNA of immune cells (“immune receptor sequencing”). But reading the DNA of immune cells does not necessarily mean we understand how the DNA encodes immune memory. Just like a German may be able to read out loud a text in French (albeit with an accent), it doesn’t mean that he understands the text.
Similarly, they wrote, the record of past encounters with antigen is encoded into the DNA in a way that we currently don’t really understand. While we do understand what each letter of the DNA means, we don’t understand how a set of letters (or words) describe the memory of previous infections. If one could decipher how past immune battles are encoded into our immunological memory, then we would be able to design more effective vaccines.
Vaccine design is difficult because we are designing vaccines for an entity – the immune system – that we don’t fully understand, Grieff and his team declared.
Recently, bioinformatics and machine learning have shown incredible success in identifying hidden patterns in very large sets of biological data (such as the DNA of immune cells). Researchers around the world have begun to combine computer science and immunology and even linguistics to decipher the language in which the immunological memory is written.
For example, if the pieces of DNA from each cell form a sentence, it may be possible to identify common words that describe previous encounters with same pathogen. Computer analyses, in conjunction with the extraction of ultra-large datasets, may help write the next Rosetta stone of immunology, which could help translate how our immune system keeps track of the myriad of pathogens it has encountered throughout our lifetime, they continued.
Computer analyses can help decipher the language of immune cells. For example, by using methods from evolutionary and mathematical ecology, scientists can now analyze how our immune system evolves over time in response to pathogen challenge and get a glimpse at how the best immune soldiers are selected. Such studies serve to analyze how the immune system chooses which immune cells are recruited to form the memory cell pool, giving us a better understanding of how to design more effective vaccines.
Research groups are also now looking into how our immune system detects foreign and potentially dangerous particles (called antigens) through the very detailed molecular resolution of immune-cell-antigen complexes at a subnanometer scale (1 million times smaller than a human hair). Such analyses seek to analyze how our immune system interacts with bacteria and viruses, allowing us to see through the immune systems’ “eyes.” These, in turn, could provide an improved understanding of how to reverse engineer effective vaccines.
Finally, given that experimental datasets are few and far between, teams have started to implement computational simulations of the immune system to test boundary conditions under which long-lasting and specific immune memory can develop. Computer simulations are becoming increasingly important in the biological sciences, as they help sift through millions of different immune scenarios thereby speeding up immunological discoveries.
Protecting our health is no easy task, the Grieff team concluded, but the body succeeds each and every day to fend off infections through the use of a very complicated immune language. Only thanks to new technologies and computer simulations are we able to read millions of such words.
Now remains the task of understanding their meaning. Extracting patterns with machine learning and language processing tools is a promising approach to pin down the natural language of our immune defense. If we are successful, the discovery and production of new vaccines can become an easier dialog.