Blog
iReceptor Plus partners leverage 3D-derived vocabulary to demonstrate predictability of antibody-antigen binding
13/04/2021
By Dr. Rahmad Akbar
Antibodies sense, neutralize and memorize foreign (viruses and bacteria) as well as domestic threats (cancer and autoimmunity). Thus antibody-antigen recognition (binding) is a central question in immunology. Accurate prediction of antibody-antigen binding along with the corresponding developability characteristics will accelerate the design and synthesis of fit-for-purpose antibody-based therapeutics.
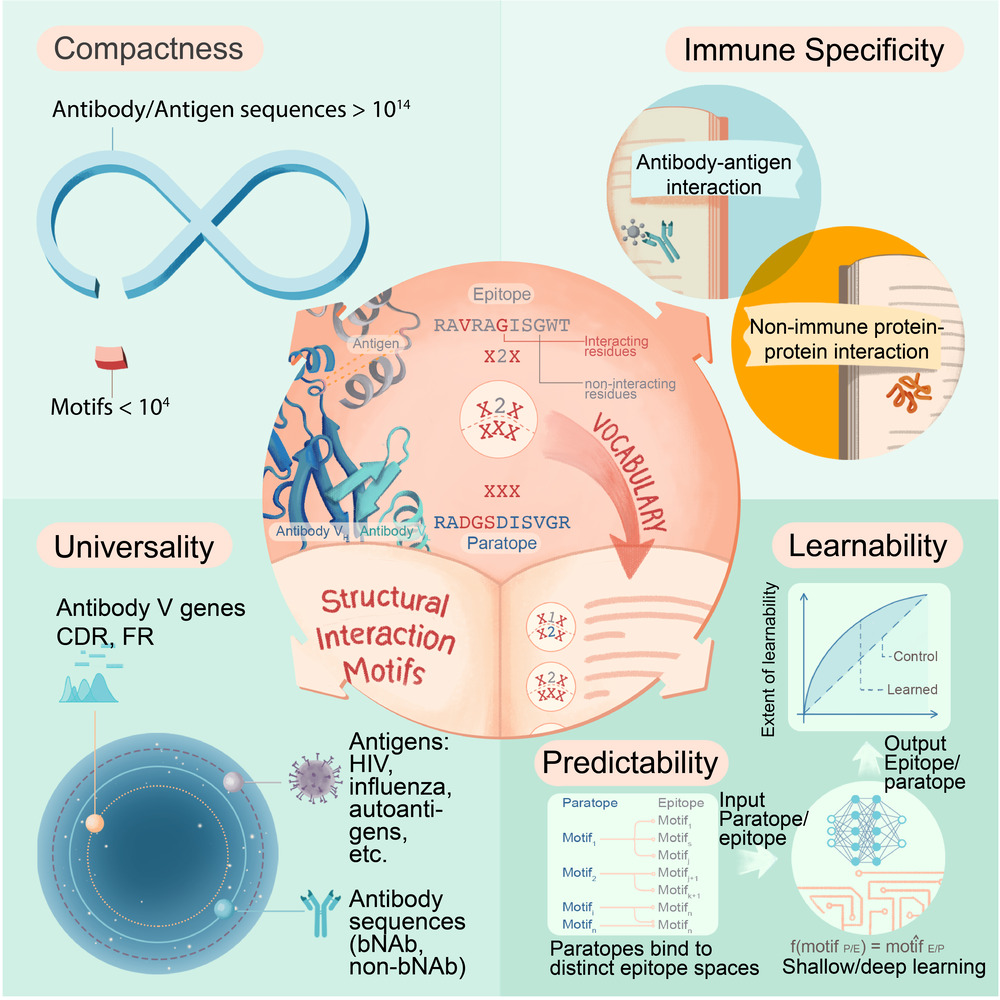
Physics teaches us that for every action there exists an equal reaction. In adaptive immunity, the vast number of possible threats necessitates an equally vast number of antibodies. Consequently, it is commonly thought that antibody-antigen binding happens at random and in unpredictable ways. However, we discovered a set of structural (3D-derived) motifs that may underlie the interactions between an antibody and an antigen. This vocabulary of motifs allowed us to demonstrate an extent of predictability therefore jettisoning the notion of complete randomness in antibody-antigen binding.
We highlight that the vocabulary we discovered is compact, immune-specific, universal, and learnable. In the future, it may pave the way for the on-demand synthesis of antibodies.
The project was supervised by Geir Kjetil Sandve (University of Oslo, UiO) and Victor Greiff (UiO, iReceptor Plus participant). Andrei Slabodkin (UiO, iReceptor Plus participant) contributed to the work.
Read the full research paper, A compact vocabulary of paratope-epitope interactions enables predictability of antibody-antigen binding, at the Cell Reports journal.
—-
The autor is a computational systems immunology scientist, at the Department of Immunology, Oslo University Hospital, University of Oslo.