Blog
iReceptor Plus partners release immuneML, a platform for machine learning analysis of adaptive immune receptor repertoires
24/03/2021
Adaptive immune receptors (T-cell and B-cell receptors, TCR and BCR, respectively) are proteins on the surface of immune cells that recognize various pathogens such as viruses and bacteria and help eliminate them. Collectively, the body’s immune receptors are called the adaptive immune receptor repertoire (AIRR).
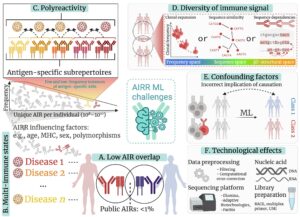
AIRRs are key targets for biomedical research because they record both past and ongoing adaptive immune responses. Understanding the immune receptor-antigen recognition process and analyzing AIRRs can help us improve the diagnosis and treatment of diseases.
In recent years, machine learning entered center stage in the biological sciences because it allows detection, recovery and re-creation of high-complexity biological information from large-scale biological data. The capacity of machine learning to identify complex discriminative sequence patterns makes it an ideal approach for AIRR-based diagnostic and therapeutic discovery.
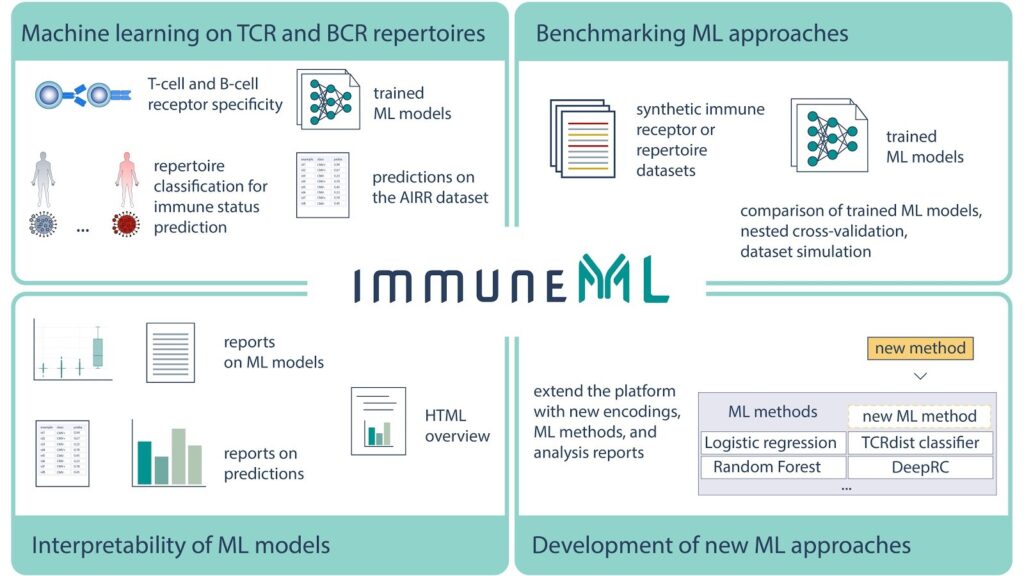
Until now, the widespread adoption of AIRR machine learning has been inhibited by a lack of reproducibility, transparency and interoperability of existing approaches. To address these concerns, as well as provide standardized workflows for development and evaluation of machine learning models on AIRR data, researchers from the University of Oslo and iReceptor Plus partners recently released immuneML, an open-source software platform for machine learning analysis of adaptive immune receptor repertoires (see also the accompanying Twitter thread). immuneML is available as a command-line tool and through an intuitive Galaxy web interface.
The manuscript describing the platform is available as a preprint on biorxiv titled “immuneML: an ecosystem for machine learning analysis of adaptive immune receptor repertoires”.
In this manuscript, the broad applicability of immuneML is showcased through three use cases:
- Reproduction of a large-scale study on immune-state prediction.
- Development, integration, and application of a novel method for antigen specificity prediction.
- Benchmarking various machine learning methods using synthetic ground truth AIRR data.
The project was supervised by Geir Kjetil Sandve (University of Oslo, UiO) and Victor Greiff (UiO, iReceptor Plus participant). Many iReceptor Plus participants contributed to immuneML: Brian Corrie, Alex Almeida Costa, Scott Christley, Lindsay Cowell, Artur Rocha, Andrei Slabodkin, Ludvig Sollid, and Gur Yaari.