Blog
Evaluating how data from different methods could – or could not – be compared
8/12/2020
By Judy Siegel-Itzkovich
Associate Prof. Encarnita Mariotti-Ferrandiz of Sorbonne University’s Immunology, Immunopathology and Immunotherapy (i3) laboratory at Pitié-Salpêtrière Hospital in Paris has headed an 18-member research team that recently published an important paper in Nature Biotechnology.
Entitled “Benchmarking of T cell receptor repertoire profiling methods reveals large systematic biases”, the article‘s data will be shared through the iReceptor Plus gateway.
Mariotti-Ferrandiz headed the team and was involved in the conception, project management and scientific and technical supervision. Prof. David Klatzmann, head of the i3 lab and another prominent iReceptor Plus team member, also contributed to the work. Pierre Barennes, who recently joined iReceptor Plus project, is a PhD fellow under Klatzmann and Mariotti-Ferrandiz supervision, who performed most of the data analysis of the study,
Monitoring the T cell receptor (TCR) repertoire in health and disease can provide key insights into adaptive immune responses, the team wrote. “But the accuracy of current TCR sequencing (TCRseq) methods is unclear.”
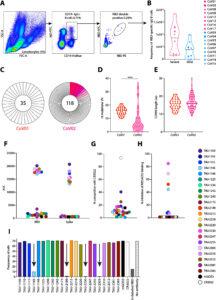
In this study, they systematically compared the results of nine commercial and academic TCRseq methods, including six rapid amplification of complementary DNA ends (RACE)-polymerase chain reaction (PCR) and three multiplex-PCR approaches, when applied to the same T cell sample.
“We found marked differences in accuracy and intra- and inter-method reproducibility for T cell receptor α (TRA) and T cell receptor β (TRB) TCR chains. Most methods showed a lower ability to capture TRA than TRB diversity. Low RNA input generated non-representative repertoires. Results from the 5′ RACE-PCR methods were consistent among themselves but differed from the RNA-based multiplex-PCR results,” they wrote.
“Using an in silico meta-repertoire generated from 108 replicates, we found that one genomic DNA-based method and two non-unique molecular identifier (UMI) RNA-based methods were more sensitive than UMI methods in detecting rare clonotypes, despite the better clonotype quantification accuracy of the latter.”
Altogether, this study, unique in its collaborative form to compare methods, revealed major differences but more importantly may allow to build sophisticated algorithms to correct intra- and maybe inter- method bias. This perspective is of utmost importance given the increasing number of publically available datasets.
Associate Prof. Mariotti-Ferrandiz’s main interest as a researcher is diversity of the immune repertoire – how it is physiologically generated and modified in pathological contexts, with a final goal to develop affordable but accurate strategies for the monitoring of the immune response as prognostic, diagnostic and prevention.
She also has experience and knowledge in several experimental models in parasitic infections (cerebral malaria), autoimmune disease (diabetes, systemic lymphoproliferative syndrome and inflammation (colitis) and in translational research with a major focus on autoimmune disorders.
Finally she supervises the development of analytical tools and modelling approaches to allow making biological sense of such complex TCR repertoire data. She gains her expertise through her successive position at Institut Pasteur (Paris, France), Riken Institute (Yokohama, Japan) and Sorbonne Université (Paris, France).
The Sorbonne Université lab is working to advance the frontiers of knowledge in Immunology and develop novel immunotherapies, with a dual reductionist and systems biology approach. Within this translational systems immunology project, the team’s focus is on tolerance in health and diseases including the role of TCR repertoire generation and variation in the physio- and pathological processes.