Blog
iReceptor Gateway utilized by researchers to track SARS-CoV-2 T-cells
3/11/2020
By Judy Siegel-Itzkovich
Following the call from the iReceptor Plus Project and the AIRR Community for increased data sharing to help overcome the COVID-19 pandemic, many COVID-19 researchers have provided data to the AIRR COVID-19 repository.
At present, some 750 million receptor sequences from 17 studies of COVID-19 patients are stored in this repository and are accessible through the iReceptor Gateway.
A major goal of iReceptor Plus is to help researchers quickly search through huge repositories of AIRR-seq data (antibody/B-cell and T-cell receptor sequences) for other data sets that can be combined to bolster or invalidate trends observed in data from a smaller number of patients. A recent study shows the power of this approach.
Researchers have used the iReceptor Gateway to pull down data from all the COVID-19 studies that have longitudinal data. One needs longitudinal data to observe temporal trends, in this case an increase in clonal diversity with time of infection. The authors used the Gateway to get more data sets that had longitudinal data – the key use of the iReceptor version 1.0, before sophisticated analysis tools are added.
“We are happy that the iReceptor Gateway was able to contribute to such an important study, and look forward to helping other researchers test hypotheses on how the adaptive immune system responds to infection, and thus contribute to the development of diagnostics and vaccines against the novel coronavirus,” said Prof. Felix Breden, Scientific Director of iReceptor Plus.
Overcoming the coronavirus pandemic will require unprecedented cooperation from the research community. This is especially true for the Adaptive Immune Receptor Repertoire Community (AIRR-C), given the vital importance of antibodies and T-cells for vaccines, diagnostics, and therapeutics in viral infection.
The AIRR-C thus has called upon its members and the wider research community to share experiences, resources, samples, and data as openly and freely as possible and to work within their respective systems to break down barriers to reach this goal, subject to the overarching directives of respect, privacy, and protection for patients and all people.
One trend observed in data from COVID-19 patients is that the number of T-cell clones that recognize and bind to SARS-Cov-2 virus increases from early to late in the infection. Such patterns are clues to how the adaptive immune system responds to the virus and can be used to monitor infections or to discover candidate molecules for therapeutics and vaccines. The ability to test this trend of course demands having data from patients who were tested multiple times during infection.
An excellent article just printed on the bioRxiv (pronounced “bio-archive”) site – a free online archive and distribution service for unpublished preprints in the life sciences that is operated by the non-profit Cold Spring Harbor Laboratory – was written by Pieter Meysman, Anna Postovskaya and colleagues at the University of Antwerp, Belgium. It was entitled “Tracking SARS-CoV-2 T cells with epitope-T-cell receptor recognition models”.
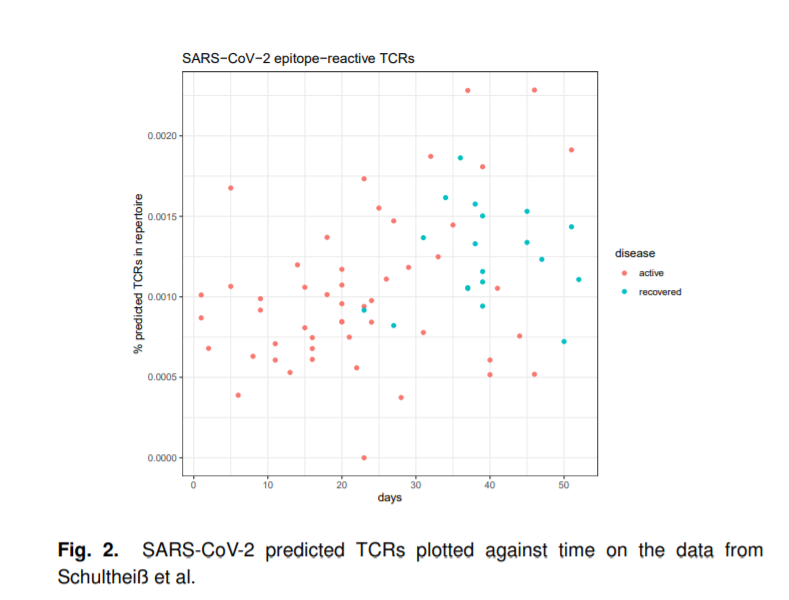
“The emergence of a novel coronavirus in 2019, termed SARS-CoV-2, has led to the most prominent global pandemic in recent history. Infection by SARS-CoV-2 manifests as COVID-19, a disease of which symptoms and severity greatly vary, and which has caused substantial loss of life all over the world,” they wrote. “Characterizing the immune response against this novel virus has become a top priority, in the hope that novel insights can lead to new treatment plans or can aid in vaccine development.”
Special attention is now being paid to T-cell response, which was already determined an important factor in long-term immunity against coronaviruses during SARS and MERS outbreaks. “Much is still not understood about the human adaptive immune response to SARS-CoV-2, the causative agent of COVID-19,” they continued.
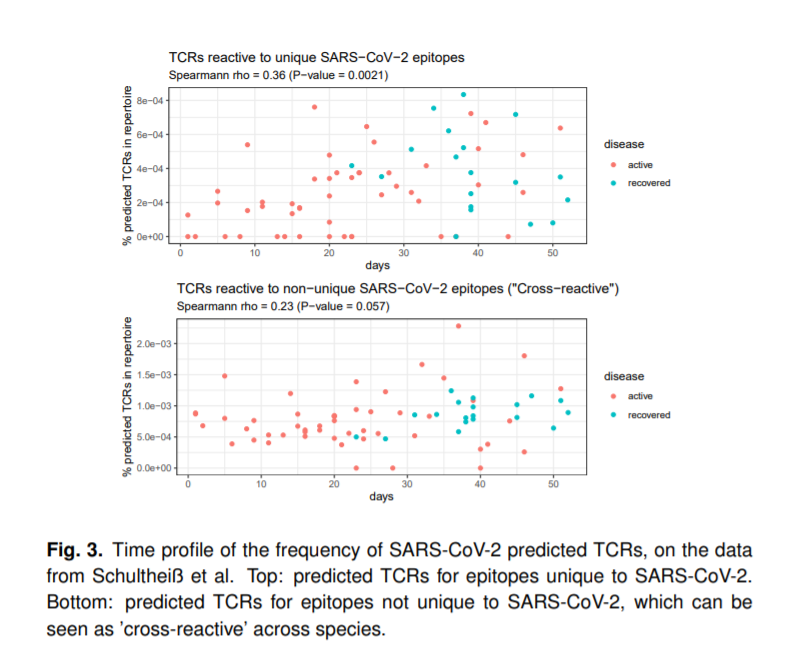
“In this paper, we demonstrate the use of machine learning to classify SARS-CoV-2 epitope specific T-cell clonotypes in T-cell receptor (TCR) sequencing data. We apply these models to public TCR data and show how they can be used to study T-cell longitudinal profiles in COVID-19 patients to characterize how the adaptive immune system reacts to the SARS-CoV-2 virus.”
Their findings confirmed prior knowledge that SARS-CoV-2 reactive T-cell diversity increases over the course of disease progression. However, “our results show a difference between those T-cells that react to epitope unique to SARS-CoV-2, which show more prominent increase, and those T-cells that react to epitopes common to other coronaviruses, which begin at a higher baseline,” they stressed.
They noted that SARS-specific T-cells have still been found in individuals 17 years later, demonstrating robust cross-reactivity against SARS-CoV-2 and suggesting the possibility of long-term protection for SARS-CoV-2. “Current findings indicate that T-cells play a key role in susceptibility to and severity of the ongoing COVID-19 pandemic as well. In particular, disease severity has been found to be linked with a sub-optimal or excessive T-cell response.”
While most of the patients present with high antibody titers frequently, they continued,” they do not provide necessary virus neutralization. Moreover, some studies have reported that around 30 percent of patients recover with very low levels of neutralizing antibodies or none at all. Most of the patients, however, developed CD8+ T-cell response, with cells predominantly showing activation signals.
Activated CD4+ T-cells were also detected in most patients, sometimes even reaching 100% in convalescent patients. Interestingly, 83% of patients were found to have CD4+ T-cells specific to the SARS-CoV-2 spike protein in one study, and the magnitude of antibody response was correlated to the CD4+ T-cell with the same protein specificity in another.”
Surprisingly, T-cells specific to the SARS-CoV-2 epitopes were even found in a high proportion of unexposed individuals, “which was later demonstrated to be due to cross-reactivity with common cold coronaviruses and might explain pre-existing protection from the SARS-CoV-2 in some individuals. Altogether those findings support a key role for T-cells in the immune defense against SARS-CoV-2 and are important considerations for vaccine development,” the authors concluded.